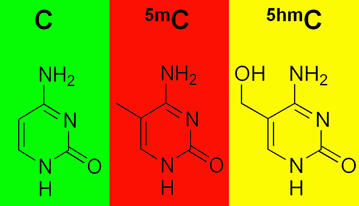
In a model of human fetal brain development, Emory researchers can see perturbations of epigenetic markers in cells derived from people with familial early-onset Alzheimer’s disease, which takes decades to appear. This suggests that in people who inherit mutations linked to early-onset Alzheimer’s, it would be possible to detect molecular changes in their brains before birth.
The results were published in the journal Cell Reports.
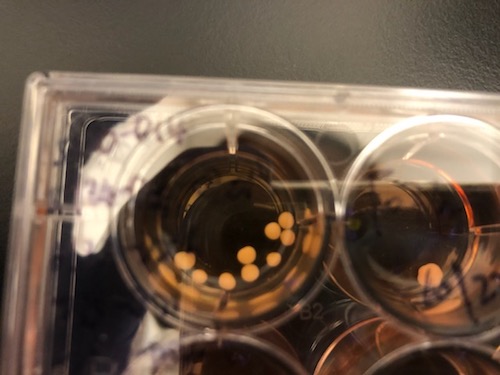
“The beauty of using organoids is that they allow us to trace back what could happen at the molecular level in early developmental stages,” says lead author Bing Yao, PhD, assistant professor of human genetics at Emory University School of Medicine. “A lot of epigenetic studies on Alzheimer’s use postmortem brains, which only represent the end point of the disease, in terms of molecular signatures.”
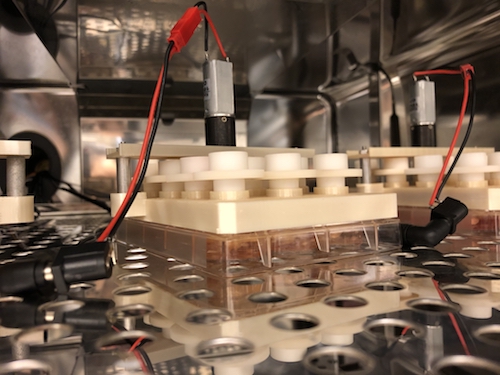
The brain organoid model allows scientists to probe human fetal brain development without poking into any babies; they have also been used to study schizophrenia, fragile X syndrome and susceptibility to Zika virus.
Co-author Zhexing Wen helped develop the model, in which human pluripotent stem cells recapitulate early stages of brain development, corresponding to 17-20 weeks after conception. The stem cell lines were obtained from both healthy donors and from people with mutations in PSEN1 or APP genes, which lead to early-onset Alzheimer’s.