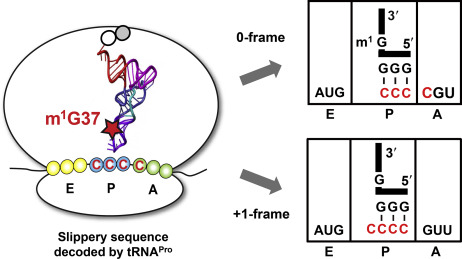
It’s a fundamental rule governing how the genetic code works. Ribosomes, the factories that assemble proteins in all types of living cells, read three letters (or nucleotides) of messenger RNA at a time.
In some instances, the ribosome can bend its rules, and read either two or four nucleotides, altering how downstream information is read. Biologists call this normally rare event ribosomal frameshifting. For an ordinary gene, the event of a frameshift turns the rest of the ensuing protein into nonsense. However, many viruses exploit frameshifting, because they can then have overlapping genes and fit more information into a limited space.
Regulated frameshifting takes place in human genes too, and understanding frameshifting is key to recent efforts to expand the genetic code. Researchers are aiming to use the process to customize proteins for industrial and pharmaceutical applications, by inserting amino acid building blocks not found in nature.
“Going back to the 1960s, when the genetic code was first revealed, there were many studies on ribosomal frameshifting, yet no-one really knows how it works on a molecular and mechanistic level,” says Christine Dunham, PhD, assistant professor of biochemistry at Emory University School of Medicine. “What we do know is that the ‘yardstick’ model that appears in a lot of textbooks, saying that the anticodon loop dictates the number of nucleotides decoded, while elegant, is probably incorrect.”
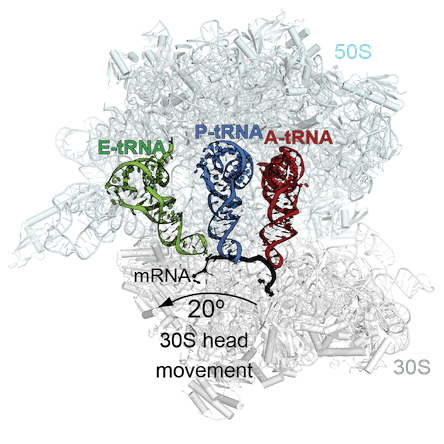
Dunham, who first studied the topic as a postdoc, and her colleagues published a paper this week in PNAS where they outline a model for how ribosomal frameshifting occurs, based on structural studies of the ribosome interacting with some of its helper machinery. Co-first authors of the paper are postdoctoral fellows Tatsuya Maehigashi, PhD and Jack Dunkle, PhD.
Read more