Efforts to produce a vaccine against HIV/AIDS have been sustained for more than a decade by a single, modest success: the RV144 clinical trial in Thailand, whose results were reported in 2009.
Now Emory, Harvard and Case Western Reserve scientists have identified a gene activity signature that may explain why the vaccine regimen in the RV144 study was protective in some individuals, while other HIV vaccine studies were not successful.
The researchers think that this signature, observed in immune cells in the blood after vaccination, could be used to design future vaccines that will have a better chance of providing protection against HIV infection.
“We may not need to take ‘shots in the dark’, when testing vaccine platforms or adjuvants for efficacy,” says senior author Rafick-Pierre Sekaly, PhD. “Instead, we can now identify adjuvants and/or vaccine regimens which more potently induce the activation of this signature.”
The results, published this week in Nature Immunology, also contain hints on a contributing factor explaining why a recent HIV vaccine study conducted in South Africa (HVTN702) did not show a protective effect. HVTN702 was designed as a follow-up to RV144, but multiple parameters were different between the Thai and South African vaccine studies, such as the demographics of the participants, the adjuvant used, and the levels and varieties of HIV circulating.
“Our findings highlight one potential mechanism which may have contributed to the muted efficacy of HVTN702,” says Sekaly, professor of pathology and laboratory medicine at Emory University School of Medicine and a Georgia Research Alliance Eminent Scholar.
This mechanism involves the choice of adjuvant, a vaccine additive that enhances immune responses. While RV144 used the adjuvant alum (aluminum hydroxide), HVTN702 used the oil-based adjuvant MF59, also found in some influenza vaccines, to stimulate higher antibody production.
“There are multiple ways that a vaccine can promote protection and some of these do not involve antibodies,” Sekaly says. “Since MF59 failed to potently induce the gene signature we found to be associated with protection, this signature could guide us to mechanisms distinct from antibodies which could trigger protection from HIV-1.”
The lead author of the Nature Immunology paper is Jeffrey Tomalka, PhD, an Instructor at Emory University School of Medicine, with co-senior author Sampa Santra, PhD, at Harvard. The research was begun while Sekaly’s lab was at Case Western Reserve, and non-human primate experiments were conducted at New England Primate Research Center and the National Institutes of Health.
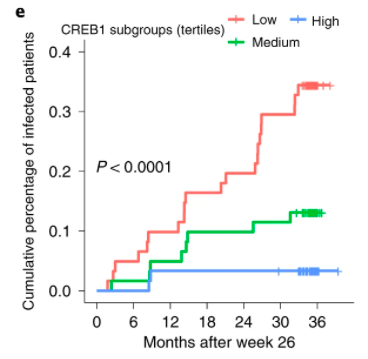
The new paper starts with studies of non-human primates immunized with ALVAC-HIV, part of the vaccine regimen in the RV144 trial. The researchers used data from those experiments to analyze samples from RV144 trial participants. In the RV144 clinical trial, volunteers who received the experimental vaccine regimen were 31 percent less likely to develop HIV infection than those who received a placebo. The gene activity signature could predict which vaccinated volunteers were likely NOT to become infected for up to 3 years. The researchers then confirmed their findings in another non-human primate vaccination study matching the RV144 regimen.
The signature of vaccine protection is not based on one gene or immune cell type. Rather, it is based on the transcription factor CREB1, a master regulator of many genes necessary for different arms of the adaptive immune system – T cells, B cells, and dendritic cells — to work together. CREB1 controls the production of several immune messengers, called cytokines and chemokines, enhances immune cells’ function, and promotes migration of immune cells to the site of vaccination.
“CREB1 is a very well-studied transcription factor, and in a sense, it was hiding in plain sight,” Tomalka says.
Previous analyses of RV144 participants’ samples had identified a type of antibody directed against part of HIV, called V1V2, that provided a protective effect. A strong CREB1 signature is correlated with V1V2 levels, but it goes beyond antibody production, Tomalka says.
“One size does not fit all when it comes to protective vaccine responses,” he says. “Focusing on a small set of targets or a specific clinical outcome, such as levels of V1V2, is likely to miss the mark in identifying the multitude of immune responses that contribute to protection.”
The research was supported by the National Institute of Allergy and Infectious Diseases (AI067854, AI100645, AI144371, P30AI060354), the Bill and Melinda Gates Foundation, and the NIH Director’s Office (Primate Centers: OD011103, OD011132).


